Network regularization in imaging-genetics
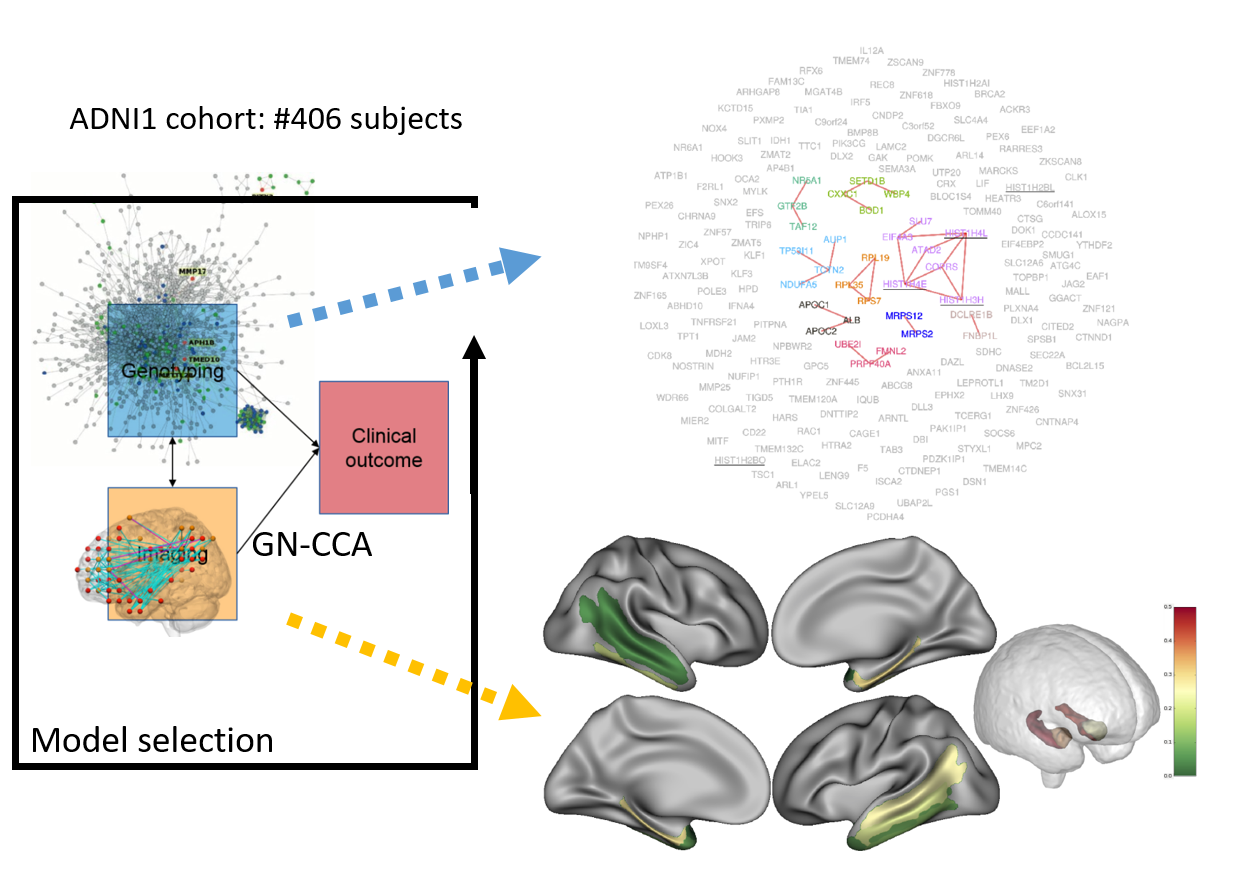
© NeuroSpin/CEA. Two components are extracted and displayed. The genes selected appear as in subgraphs of the protein-protein iteraction graph. Regions selected are spatially coherent.
In this work, we demonstrate the interest GN-CCA, a method for imaging-genetics data integration with structured prior knowledge. We used the ADNI dataset. The genetic block is transformed in the gene mutation burden matrix (gene x subject matrix). The mean cortical thickness is estimated in Destrieux ROI set (roi x subject matrix). Those two blocks were analyzed using a customized CCA exploratory approach that accommodate the intra block correlation structure to yield one component for each block. In those components the weight of two genes that are linked in the protein-protein interaction graph are forced to be similar. The weights of patially close regions are constrained similarly. Hyperparameters of this approach are chosen using model selection framework that use the components estimated by the regularized CCA as predictors. The components obtained not only display good performance, but they are also easy to interpret in terms of biology.
Guigui, N., Philippe, C., Gloaguen, A., Karkar, S., Guillemot, V., Löfstedt, T., & Frouin, V. (2009). Network regularization in imaging genetics improves prediction performances and model interpretability on Alzheimers’s disease. In ISBI 2019 - Proceedings of the IEEE International Symposium on Biomedical Imaging. Venice, Italy. Retrieved from https://hal-cea.archives-ouvertes.fr/cea-02016625